Note
Click here to download the full example code
Plot MEG inverse solution¶
Data were computed using mne-python (http://martinos.org/mne)
Set up some useful variables and make the plot.
# define subject, surface and hemisphere(s) to plot:
subject_id, surf = 'fsaverage', 'inflated'
hemi = 'lh'
# create Brain object for visualization
brain = Brain(subject_id, hemi, surf, size=(400, 400), background='w',
interaction='terrain', cortex='bone', units='m')
# label for time annotation in milliseconds
def time_label(t):
return 'time=%0.2f ms' % (t * 1e3)
# Read MNE dSPM inverse solution and plot
for hemi in ['lh']: # , 'rh']:
stc_fname = os.path.join('example_data', 'meg_source_estimate-' +
hemi + '.stc')
stc = read_stc(stc_fname)
# data and vertices for which the data is defined
data = stc['data']
vertices = stc['vertices']
# time points (in seconds)
time = np.linspace(stc['tmin'], stc['tmin'] + data.shape[1] * stc['tstep'],
data.shape[1], endpoint=False)
# colormap to use
colormap = 'hot'
# add data and set the initial time displayed to 100 ms,
# plotted using the nearest relevant colors
brain.add_data(data, colormap=colormap, vertices=vertices,
smoothing_steps='nearest', time=time, time_label=time_label,
hemi=hemi, initial_time=0.1, verbose=False)
# scale colormap
brain.scale_data_colormap(fmin=13, fmid=18, fmax=22, transparent=True,
verbose=False)
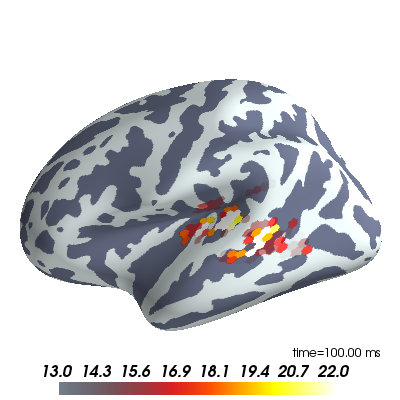
To change the time displayed to 80 ms uncomment this line:
# brain.set_time(0.08)
uncomment these lines to use the interactive TimeViewer GUI
# from surfer import TimeViewer
# viewer = TimeViewer(brain)
Total running time of the script: ( 0 minutes 0.734 seconds)
